New paper from our laboratory
Kudoh, A., Megonigal, J. P., Langley, J. A., Noyce, G. L., Sakai, T., & Whigham, D. F. (2023). Reproductive Responses to Increased Shoot Density and Global Change Drivers in a Widespread Clonal Wetland Species, Schoenoplectus americanus. Estuaries and Coasts. https://doi.org/10.1007/s12237-023-01249-z
Abstract
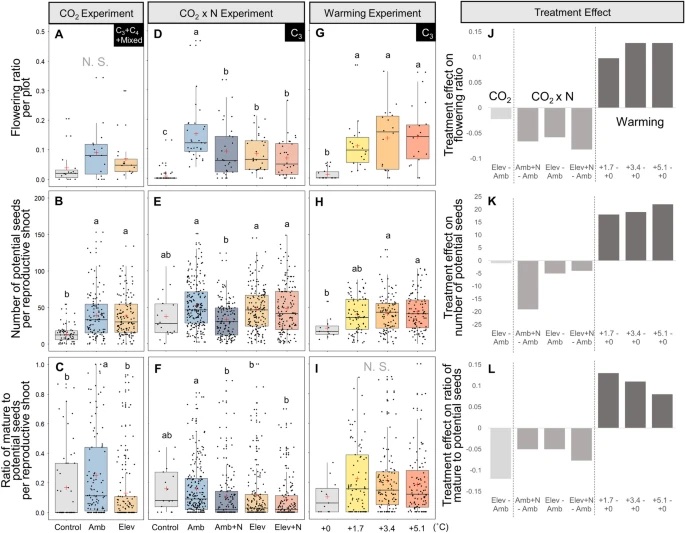
The expansion of many wetland species is a function of both clonal propagation and sexual reproduction. The production of ramets through clonal propagation enables plants to move and occupy space near parent ramets, while seeds produced by sexual reproduction enable species to disperse and colonize open or disturbed sites both near and far from parents. The balance between clonal propagation and sexual reproduction is known to vary with plant density but few studies have focused on reproductive allocation with density changes in response to global climate change. Schoenoplectus americanus is a widespread clonal wetland species in North America and a dominant species in Chesapeake Bay brackish tidal wetlands. Long-term experiments on responses of S. americanus to global change provided the opportunity to compare the two modes of propagation under different treatments. Seed production increased with increasing shoot density, supporting the hypothesis that factors causing increased clonal reproduction (e.g., higher shoot density) stimulate sexual reproduction and dispersal of genets. The increase in allocation to sexual reproduction was mainly the result of an increase in the number of ramets that flowered and not an increase in the number of seeds per reproductive shoot, or the ratio between the number of flowers produced per inflorescence and the number of flowers that developed into seeds. Seed production increased in response to increasing temperatures and decreased or did not change in response to increased CO2 or nitrogen. Results from this comparative study demonstrate that plant responses to global change treatments affect resource allocation and can alter the ability of species to produce seeds.